| ZFP36L1 |
|---|
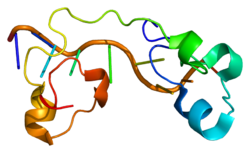 |
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
1W0V, 1W0W |
|
|
| Identifiers |
|---|
| Aliases | ZFP36L1, BRF1, Berg36, ERF-1, ERF1, RNF162B, TIS11B, cMG1, ZFP36 ring finger protein-like 1, ZFP36 ring finger protein like 1 |
|---|
| External IDs | OMIM: 601064; MGI: 107946; HomoloGene: 31276; GeneCards: ZFP36L1; OMA:ZFP36L1 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 14 (human)[1] |
|---|
| | Band | 14q24.1 | Start | 68,787,660 bp[1] |
|---|
| End | 68,796,253 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 12 (mouse)[2] |
|---|
| | Band | 12|12 C3 | Start | 80,154,528 bp[2] |
|---|
| End | 80,159,787 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - mucosa of paranasal sinus
- canal of the cervix
- gastric mucosa
- right ovary
- ectocervix
- nipple
- left ovary
- urethra
- body of uterus
- right uterine tube
|
| | Top expressed in | - corneal stroma
- genital tubercle
- ventricular zone
- lactiferous gland
- lip
- yolk sac
- esophagus
- cardiac muscle tissue of left ventricle
- superior surface of tongue
- atrium
|
| | More reference expression data |
|
|---|
| BioGPS | 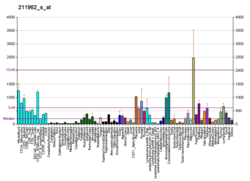
 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - DNA binding
- DNA-binding transcription factor activity
- metal ion binding
- protein binding
- mRNA binding
- 14-3-3 protein binding
- RNA binding
- mRNA 3'-UTR AU-rich region binding
- DNA-binding transcription factor activity, RNA polymerase II-specific
- mRNA 3'-UTR binding
| | Cellular component | - cytoplasm
- nucleus
- cytosol
- P-body
- ribonucleoprotein complex
| | Biological process | - spongiotrophoblast layer development
- mRNA catabolic process
- multicellular organism growth
- embryonic organ development
- chorio-allantoic fusion
- vasculogenesis
- proepicardium development
- heart development
- nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay
- neural tube development
- cell population proliferation
- regulation of translation
- apoptotic process
- T cell differentiation in thymus
- regulation of gene expression
- negative regulation of erythrocyte differentiation
- cellular response to insulin stimulus
- cellular response to phorbol 13-acetate 12-myristate
- ERK1 and ERK2 cascade
- positive regulation of intracellular mRNA localization
- cellular response to glucocorticoid stimulus
- cellular response to transforming growth factor beta stimulus
- regulation of keratinocyte differentiation
- response to wounding
- positive regulation of monocyte differentiation
- regulation of transcription, DNA-templated
- cellular response to hypoxia
- regulation of keratinocyte apoptotic process
- regulation of keratinocyte proliferation
- phosphatidylinositol 3-kinase signaling
- cellular response to tumor necrosis factor
- cellular response to epidermal growth factor stimulus
- regulation of mRNA 3'-end processing
- MAPK cascade
- mRNA processing
- multicellular organism development
- nuclear-transcribed mRNA catabolic process, deadenylation-independent decay
- p38MAPK cascade
- regulation of mRNA stability
- protein kinase B signaling
- cellular response to fibroblast growth factor stimulus
- regulation of B cell differentiation
- positive regulation of fat cell differentiation
- regulation of myoblast differentiation
- mesendoderm development
- mRNA transport
- 3'-UTR-mediated mRNA destabilization
- cellular response to cAMP
- cellular response to peptide hormone stimulus
- cellular response to salt stress
- regulation of stem cell proliferation
- cellular response to raffinose
- positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay
- negative regulation of mitotic cell cycle phase transition
- transport
- regulation of transcription by RNA polymerase II
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_004926
NM_001244698
NM_001244701 |
| |
|---|
| RefSeq (protein) | |
|---|
NP_001231627
NP_001231630
NP_004917 |
| |
|---|
| Location (UCSC) | Chr 14: 68.79 – 68.8 Mb | Chr 12: 80.15 – 80.16 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|

 1rgo: Structural Basis for Recognition of the mRNA Class II AU-Rich Element by the Tandem Zinc Finger Domain of TIS11d
1rgo: Structural Basis for Recognition of the mRNA Class II AU-Rich Element by the Tandem Zinc Finger Domain of TIS11d


















 ZFP36L1
ZFP36L1